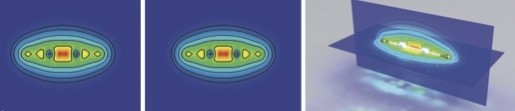
Atomistic simulations should always be compared to experimental results (a theme which is developed in Chapter 18, Comparison to Experiment). These comparisons allow us to test the reliability of the simulations, and to learn more about the experiments: ultimately, we should have enough confidence in the simulations to interpret and explain experimental results.
A recent paper in Phys. Rev. Lett.[1] takes this approach for a large defect in bulk silicon induced by ion implantation. (Ion implantation is one standard route to dope silicon and germanium for electronic devices, and is normally followed by annealing.) Most importantly, the experimental technique used, high-angle annular dark field scanning transmission electron microscopy (HAADF STEM), was compared to modelling without any processing or structure fitting. Normally, high resolution TEM requires some image processing.
In the modelling, the defect was induced by creating a series of silicon interstitials and local re-arrangements of atoms (which they refer to as bond defects). The length of the defect was chosen to match experiment, and the system was simulated using the Tersoff potential, annealed at 1400K. The excess heat generated by relaxations of the high energy starting structure was removed by velocity rescaling (a more sophisticated thermostat could have been used, but may not have been needed). The orientation of the defect, along {311}, is set by the initial structure, and after 2ns of annealing the defect had transformed into a series of five-, seven- and eight-membered rings along with chains of interstitials.
In the paper, this structure is compared to an experimental image, and the agreement is excellent: the two structures are the same, even down to slight differences in bond lengths and angles in the rings of the defects (lengths agree to within 0.5 Angstroms, angles to within a few degrees). These comparisons required some image processing, but overall the two approaches agree to a remarkable degree, and validate each other. However, it’s important to note that analysis of only one defect is presented, and there are few details given of how changes in the initial conditions affected the final structure found by MD. We don’t know how agreement would change with defect length, for instance.
What are the lessons to be learnt ? Certainly that molecular dynamics with relatively simple forcefields is capable of quantitative, predictive accuracy when the problem is chosen carefully. The bonding and angles in this problem are sufficiently close to bulk-like that Tersoff works well: the problem lies within the domain where the potential was fitted. A relatively modest simulation, using 4,800 atoms, and performing simple MD was effective. Notice what was compared: not total energies (which many DFT practitioners rely on) but structural data. This is appropriate here, particularly as the system generated was not necessarily the thermodynamic minimum. It’s vital that we move away from the knee-jerk assumption that lowest energy in simulation is what should be seen in experiment (see Section 18.2.1 for vivid illustration of this).
In the slightly longer term, it will be interesting to see if large-scale DFT techniques such as linear scaling DFT (see http://www.order-n.org/ and a recent review[2] for more details) will be able to contribute to this area, and generate more quantitative information.
[1] Phys. Rev. Lett. 110, 166102 (2013) DOI:10.1103/PhysRevLett.110.166102